One of the primary focuses of our lab is a group of bacteria known as rhizobia, with a particular emphasis on the rhizobium Sinorhizobium meliloti. Rhizobia are a fascinating group of soil-dwelling Proteobacteria able to enter into endosymbiotic relationships with leguminous plants, during which they inhabit a plant organ known as a nodule. Here, the rhizobia convert atmospheric nitrogen gas into ammonium that is transferred to the plant in exchange for a source of carbon, thereby providing legumes a distinct ability to thrive in nitrogen-deficient soils. This process of symbiotic nitrogen fixation is of significant scientific interest due to its tremendous ecological and agricultural significance. In addition to being a leading model for the study of symbiotic nitrogen fixation, S. meliloti is a wonderful system to examine bacterial metabolism and genome evolution, among other areas. Below, some of the main topics of interest within our general rhizobium program are outlined, and we are always excited to explore new directions not listed below.
Genetics and metabolism of nodulation and nitrogen fixation
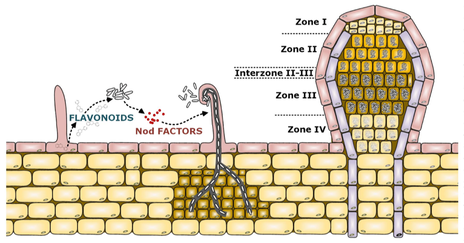
A schematic representation of the various stages of rhizobium - legume symbioses. Shown is the initial exchange of signals (left), growth of the bacterium along an infection thread (middle), and an indeterminate nodule (right). Figure prepared by Dr. Maryam Zamani, and adapted from [21].
Symbioses between rhizobia and legumes are exceedingly complex. Following an exchange of signals (Nod factors produced by the rhizobia and flavonoids produced by the plants) [10-12], the rhizobia penetrate legume root hairs where they replicate within a growing infection thread [13]. Upon reaching the cortical layer, the rhizobia are released from the infection thread and enter nodule cells through endocytosis [13-15]. The process of endocytosis results in the bacteria, now referred to as bacteroids, being surrounded by a plant derived membrane known as the peribacteroid membrane. The plant and bacterial cells undergo a dramatic developmental progression involving cell size enlargement, endoreduplication, and major transcriptional, morphological, and metabolic changes [16-18]. Finally, the bacteroids convert N2 gas into ammonia, which is provided to the plant in exchange for a source of carbon (C4-dicarboxylates - malate, succinate, fumarate) [19]. The developmental stages are spatially separated across distinct zone in nodules that have an indeterminate structure [18, 20], such as those formed by Medicago species. Despite that symbiotic nitrogen fixation has been well-studied [21], we remain far form a comprehensive understanding of this interaction. In our work, we aim to:
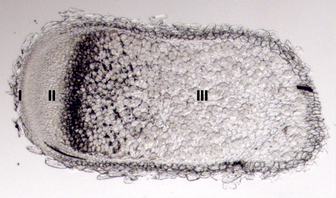

A longitudinal cross-section of a pea root nodule stained with propidium iodine to visualize starch granules. The far left is the nodule apical meristem (zone I), followed to the right by the infection and differentiation zone (zone II), and the nitrogen fixation zone (zone III). The darkly stained region between zone II and zone III is known as the interzone II-II. Figure adapted from [30].
- Develop and deploy a metabolic modelling framework to study the metabolism in each of the nodule zones. Symbiotic nitrogen fixation is predicated on a metabolic integration of the two partners [19]. Therefore, we are using genome-scale metabolic network reconstruction and constraint-based modelling to generate an in silico picture of the metabolism of a legume nodule in collaboration with Dr. Mengoni and Dr. Fondi at the University of Florence (Italy). We will use the model to generate hypotheses for experimental validation and to address questions difficult to study experimentally.
- Dissect the genetic requirements for rhizobia to effectively nodulate legumes using novel genetic tools. Most knowledge of the symbiosis related to the ultimate stage of nitrogen fixation; much less is understood about what occurs during the preceding developmental progression partly due to experimental difficulties. While the metabolic modelling strategy will help, experimental tools are required for us to fully tackle this knowledge gap. We are interested in developing new genetic tools to address this problem, and employing them to evaluate nodule zone-specific genetic requirements.
- Uncover global regulatory mechanisms controlling the differentiation process. We hope to identify general regulatory mechanisms in rhizobia that control the process of differentiation using new technologies.
- Interrogate the role of the BacA protein during nodulation and nitrogen fixation. The BacA protein plays an indispensable role during symbiosis with certain legume species [17, 22] and may function as a host-compatibility factor [23]. BacA is also found in pathogens and can play a role in infection [24]. However, the precise function of BacA remains unknown. We maintain an interest in characterizing this protein and unearthing its function.
Metabolism, and its regulation, in prokaryotic organisms
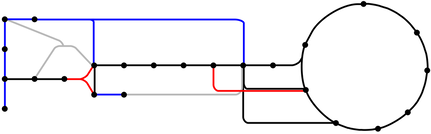
A map of the central carbon metabolic pathways of S. meliloti. Lines represent reactions and dots are metabolites. The genes associated with reactions in blue and red are repressed and induced, respectively, by the transcriptional regulator PckR. In grey are incomplete representations of pathways. Figure adapted from [28].
Metabolism - the interconversion of chemical compounds - is the cornerstone of all biological life. Organisms would not survive without the ability to degrade food, synthesize cellular building blocks, or generate stress-protective molecules. However, it is crucial that the the expression and activity of metabolic proteins is tightly regulated in response to changing environmental conditions. Rhizobia are fantastic models to study metabolism and its regulation in bacteria. The rhizobia have a life-style that involves growth in soil environments, colonization of plant roots and root-associated soils (the rhizosphere), and terminal differentiation within legume nodules followed by nitrogen fixation. Likely driven in part by their adaptation to diverse niches, rhizobia have large genomes encoding a broad metabolic repertoire [1, 2]. For example, the model strain Sinorhizobium meiloti Rm1021 can catabolize over 70 carbon sources and 50 nitrogen sources [3, 4], and its large 6.7 Mb genome encodes a whopping 200 ABC-type transport proteins [5, 6]. In our work, we aim to:
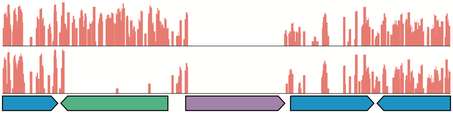
A visual illustration of two Tn-seq datasets. Each of the upper two tracks represent a Tn-seq dataset for S. meliloti when grown in different nutritional conditions [29]. Boxes along the bottom represent protein coding regions. The red bars indicate the location and abundance of transposons in a bacterial population. The blue genes are non-essential, the purple gene is essential, and the green gene is conditionally essential. Figure adapted from [21].
- Develop experimental and computational tools to support the characterization of metabolic and regulatory processes through an integrative wet-lab and dry-lab approach. This includes generating improved protocols and new variants of the functional genomics tool transposon-sequencing (Tn-seq) [7], as well as expanding our toolbox for integrating transposon-sequencing data with metabolic network reconstruction and constraint-based modelling [8].
- Generate unprecedented insight into the metabolic and regulatory networks of S. meliloti. We will evaluate the adaptation of S. meliloti to a broad range of nutritional and stress conditions using a combination of functional genomics, transcriptomics, proteomics, and metabolic modelling. These data sets will provide fundamental knowledge that will allow us to: i) generate high-quality functional and regulatory gene networks, ii) evaluate the relationships between changes in gene expression and fitness contribution, iii) functionally annotate some of the 30% of bacterial genes that continue to have no known function [9], and iv) characterize novel regulators that may be valuable in synthetic regulatory circuit.
- Analyze the evolution of metabolism in the rhizobia. Through national and inter-national collaborations, we are using Tn-seq and metabolic modelling to study the metabolism of several rhizobium species. These investigations will provide a valuable compendium to examine the common and lineage-specific metabolism of these diverse bacteria.
Taxonomy of rhizobia
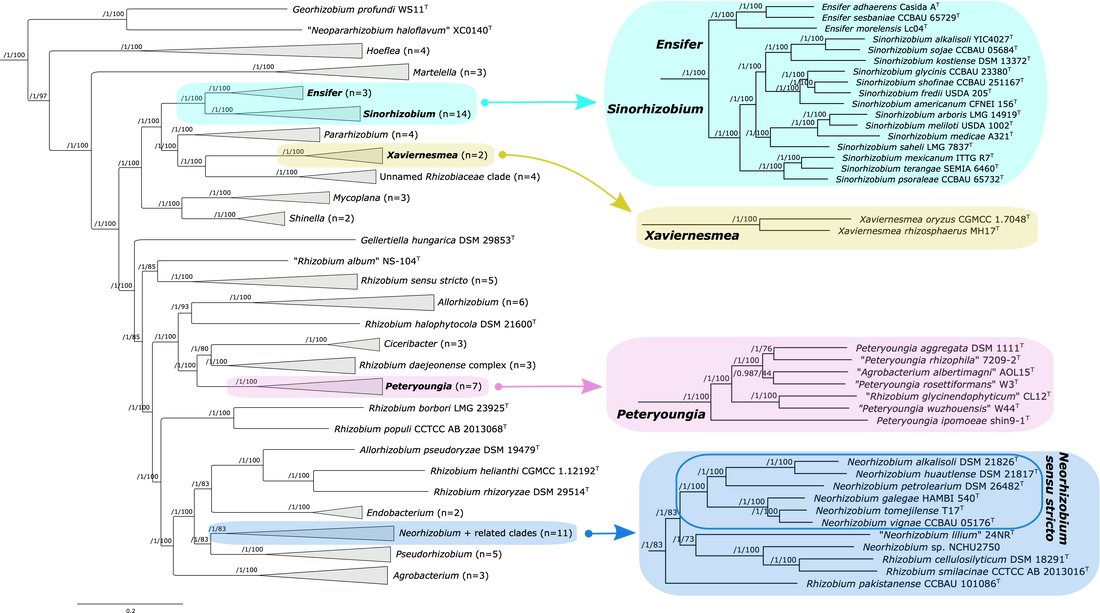
A phylogeny of the family Rhizobiacea. Figure taken from [31].
Taxonomically, rhizobia can be sub-divided into the alpha-rhizobia (found within the class Alphaproteobacteria) and beta-rhizobia (found within the class Betaproteobacteria). The alpha-rhizobia are polyphyletic and can be found in several genera throughout the order Hyphomicrobiales. Unfortunately, the taxonomy of clades containing rhizobia is "messy", in that species are often assigned to the wrong genera, and likewise, some genera are assigned to the wrong family. It is therefore necessary to fix the taxonomy of clades containing rhizobia to ensure consistent and accurate classification of new strains, and to also develop common taxonomic frameworks for use by the community. We have begun tackling this problem through development of a genome-based taxonomic framework for genus delimitation within the family Rhizobiaceae [31], but more work needs to be done. We are interested in continuing to refine the taxonomy of rhizobia using a genome-based approach, as well as defining new species and genera for publicly available strains and for newly isolated strains in our collection.
Function and evolution of multipartite genomes in bacteria
When thinking of what a bacterial genome looks like, most people are likely to imagine the structure found in Escherichia coli: one circular chromosome, possibly with some plasmids. Indeed, most bacteria have a genome that follows this architecture. However, about 10% of bacterial genomes that are split between multiple large DNA molecules (i.e., DNA replicons), which are referred to as multipartite genomes or divided genomes [25]. These large replicons include megaplasmids (essentially large plasmids) and chromids, which contain several chromosome-like feature and appear to have arose from megaplasmids [25, 26]. The multipartite genome structure is most commonly found in the Proteobacteria, and appear to be particularly prevalent in many symbionts (including rhizobia! [1]), as well as many pathogenic organisms (e.g., Burkholderia, Vibrio, and Brucella). We remain broadly interested in examining the evolution and the function of the multipartite genome structure, and investigating the consequences of the complete removal of DNA replicons from a genome [4, 27].
Funding
Support for this research has been provided by the Natural Sciences and Engineering Research Council of Canada (NSERC), the Joint Genome Institute (JGI), Mitacs, and Queen's University.
[1] MacLean AM, Finan TM, Sadowsky MJ. (2007) Genomes of the symbiotic nitrogen-fixing bacteria of legumes. Plant Physiology. 144(2): 615-622. HTML
[2] Poole P, Ramachandran V, Terpolilli J. (2018) Rhizobia: from saprophytes to endosymbionts. Nature Reviews Microbiology. 16(5) 291-303. HTML
[3] Biondi EG, Tatti E, Comparini D, Giuntini E, Mocali S, Giovannetti L, Bazzicalupo M, Mengoni A, Viti C. (2009) Metabolic capacity of Sinorhizobium (Enifer) meliloti strains as determined by phenotype microarray analysis. Applied and Environmental Microbiology. 75(16): 5396-5404. HTML
[4] diCenzo GC, MacLean AM, Milunovic B, Golding GB, Finan TM. (2014) Examination of prokaryotic multipartite genome evolution through experimental genome reduction. PLOS Genetics. 10(10): e1004742. HTML
[5] Galibert F, Finan TM, Long SR, Puhler A, ..., Batut J. (2001). The composite genome of the legume symbiont Sinorhizobium meliloti. Science. 293(5530): 668-672. HTML
[6] Mauchline TH, Fowler JE, East AK, Sartor AL, Zaheer R, hosie AH, Poole PS, Finan TM. (2006) Mapping the Sinorhizobium meliloti 1021 solute-binding protein-dependent transportome. Proceedings of the National Academy of Science of the United States of America. 103(47): 17933-17938. HTML
[7] Van Opijnen T, Camilli A. (2013) Transposon insertion sequencing: a new tool for systems-level analysis of microorganisms. Nature Reviews Microbiology. 11(7) 435-442. HTML
[8] diCenzo GC, Mengoni A, Fondi M. (2019) Tn-Core: A toolbox for integrating Tn-seq gene essentiality data and constraint-based metabolic modeling. ACS Synthetic Biology. 8(1): 158-169. HTML
[9] Hanson AD, Pribat A, Waller JC, de Crécy-lagard V. (2009) 'Unknown' proteins and 'orphan' enzymes: the missing half of the engineering parts list - and how to find it. Biochemistry Journal. 425(1): 1-11. HTML
[10] Geurts R, Bisseling T. (2002) Rhizobium Nod factor perception and signalling. The Plant Cell. 14(Suppl 1): s239-249. HTML
[11] Oldroyd GED. (2013) Speak, friend, and enter: signalling systems that promote beneficial symbiotic associations in plants. Nature Reviews Microbiology. 11(4): 252-263. HTML
[12] Long SR. (2001) Genes and signals in the Rhizobium-legume symbiosis. Plant Physiology. 125(1) 69-71. HTML
[13] Jones KM, Kobayashi H, Davies BW, Taga ME, Walker GC. (2007) How rhizobial symbionts invade plants: the Sinorhizobium-Medicago model. Nature Reviews Microbiology. 5(8): 619-833. HTML
[14] Oldroyd GE, Murray JD, Poole PS, Downie JA. (2011) The rules of engagement in the legume-rhizobial symbiosis. Annual Review of Genetics. 45: 119-144. HTML
[15] Brewin NJ. (2004) Plant cell wall remodelling in the rhizobium-legume symbiosis. Critical Reviews in Plant Sciences. 23(4): 293-316. HTML
[16] Kereszt A, Mergaert P, Kondorosi É. (2011) Bacteroid develoment in legume nodules: evolution of mutual benefit or of sacrificial victims? Molecular Plant-Microbe Interactions. 24(11): 1300-1309. HTML
[17] Haag AF, Arnold MFF, Myka KK, Kersher B, Dall'Angelo S, Zanda M, Mergaert P, Ferguson GP. (2013) Molecular insights into bacteroid development during Rhizobium-legume symbiosis. FEMS Microbiology Reviews. 37(3): 364-383. HTML
[18] Roux B, Rodde N, Jardinaud MG, Timmers T, Sauviac L, Cottret L, Carrère S, Sallet E, Courcelle E, Moreau S, Debellé F, Capela D, de Carvalho-Niebel F, Gouzy J, Bruand C, Gamas P. 2014. An integrated analysis of plant and bacterial gene expression in symbiotic root nodules using laser-capture microdissection coupled to RNA sequencing. The Plant Journal. 77(6): 827-837. HTML
[19] Udvardi M, Poole PS. (2013) Transport and metabolism in legume-rhizobia symbioses. Annual Review of Plant Biology. 64: 781-805. HTML
[20] Vasse J, de Billy F, Camut S, Truchet G. (1990) Correlation between ultrastructural differentiation of bacteroids and nitrogen fixation in alfalfa nodules. Journal of Bacteriology. 172(8): 4295-4306. HTML
[21] diCenzo GC, Zamani M, Checcucci A, Fondi M, Griffitts JS, Finan TM, Mengoni A. (2019) Multi-disciplinary approaches for studying rhizobium – legume symbioses. Canadian Journal of Microbiology. 65: 1-33. HTML
[22] Guefrachi I, Verly C, Kondorisi É, Alunni B, Mergaert P. (2015) Role of the bacterial BacA ABC-transporter in chronic infection of nodule cells by Rhizobium bacteria. In Biological Nitrogen Fixation. Edited by F. J. de Bruin. Wiley Blackwell, Hobokon, New Jersy, USA. HTML
[23] diCenzo GC, Zamani M, Ludwig H, Finan TM. (2017) Heterologous complementation reveals a specialized role for BacA in the Medicago – Sinorhizobium meliloti symbiosis. Molecular Plant-Microbe Interactions. 30(4): 312-324. HTML
[24] Domenech P, Kobayashi H, LeVier K, Walker GC, Barry CE 3rd. (2009) BacA, and ABC transporter involved in maintenance of chronic murine infections with Mycobacterium tuberculosis. Journal of Bacteriology. 191(2): 477-485. HTML
[25] diCenzo GC, Finan TM. (2017) The divided bacterial genome: structure, function, and evolution. Microbiology and Molecular Biology Reviews. 81(3): e00019-17. HTML
[26] Harrison PW, Lower RP, Kim NK, Young JP (2010) Introducing the bacterial 'chromid': not a chromosome, not a plasmid. Trends in Microbiology. 18(4): 141-148. HTML
[27] Oresnik IJ, Liu SL, Yost CK, Hynes MF. (2000) Megaplasmid pRme2011a of Sinorhizobium meliloti is not required for viability. Journal of Bacteriology. 182(12): 3582-2586. HTML
[28] diCenzo GC, Muhammed Z, Østerås M, O’Brien SAP, Finan TM. (2017) PckR is a key regulator of the glycolytic and gluconeogenic central metabolic pathways in Sinorhizobium meliloti. Genetics. 207(3): 961-974. HTML
[29] diCenzo GC, Benedict A, Fondi M, Walker G, Finan TM, Mengoni A, Griffitts JS. (2018) Robustness encoded across essential and accessory replicons of the ecologically versatile bacterium Sinorhizobium meliloti. PLOS Genetics. 14(4): e1007357. HTML
[30] Mitsch MJ, diCenzo GC, Cowie A, Finan TM. (2018) Succinate transport is not essential for symbiotic nitrogen fixation by Sinorhizobium meliloti nor Rhizobium leguminosarum. Applied and Environmental Microbiology. 84(1): e01561-17. HTML
[31] Kuzmanović N, Fagorzi C, Mengoni A, Lassalle F, diCenzo GC. (2022) Taxonomy of Rhizobiaceae revisited: proposal of a new framework for genus delimitation. International Journal of Systematic and Evolutionary Microbiology. 72(3): 005243. HTML
[2] Poole P, Ramachandran V, Terpolilli J. (2018) Rhizobia: from saprophytes to endosymbionts. Nature Reviews Microbiology. 16(5) 291-303. HTML
[3] Biondi EG, Tatti E, Comparini D, Giuntini E, Mocali S, Giovannetti L, Bazzicalupo M, Mengoni A, Viti C. (2009) Metabolic capacity of Sinorhizobium (Enifer) meliloti strains as determined by phenotype microarray analysis. Applied and Environmental Microbiology. 75(16): 5396-5404. HTML
[4] diCenzo GC, MacLean AM, Milunovic B, Golding GB, Finan TM. (2014) Examination of prokaryotic multipartite genome evolution through experimental genome reduction. PLOS Genetics. 10(10): e1004742. HTML
[5] Galibert F, Finan TM, Long SR, Puhler A, ..., Batut J. (2001). The composite genome of the legume symbiont Sinorhizobium meliloti. Science. 293(5530): 668-672. HTML
[6] Mauchline TH, Fowler JE, East AK, Sartor AL, Zaheer R, hosie AH, Poole PS, Finan TM. (2006) Mapping the Sinorhizobium meliloti 1021 solute-binding protein-dependent transportome. Proceedings of the National Academy of Science of the United States of America. 103(47): 17933-17938. HTML
[7] Van Opijnen T, Camilli A. (2013) Transposon insertion sequencing: a new tool for systems-level analysis of microorganisms. Nature Reviews Microbiology. 11(7) 435-442. HTML
[8] diCenzo GC, Mengoni A, Fondi M. (2019) Tn-Core: A toolbox for integrating Tn-seq gene essentiality data and constraint-based metabolic modeling. ACS Synthetic Biology. 8(1): 158-169. HTML
[9] Hanson AD, Pribat A, Waller JC, de Crécy-lagard V. (2009) 'Unknown' proteins and 'orphan' enzymes: the missing half of the engineering parts list - and how to find it. Biochemistry Journal. 425(1): 1-11. HTML
[10] Geurts R, Bisseling T. (2002) Rhizobium Nod factor perception and signalling. The Plant Cell. 14(Suppl 1): s239-249. HTML
[11] Oldroyd GED. (2013) Speak, friend, and enter: signalling systems that promote beneficial symbiotic associations in plants. Nature Reviews Microbiology. 11(4): 252-263. HTML
[12] Long SR. (2001) Genes and signals in the Rhizobium-legume symbiosis. Plant Physiology. 125(1) 69-71. HTML
[13] Jones KM, Kobayashi H, Davies BW, Taga ME, Walker GC. (2007) How rhizobial symbionts invade plants: the Sinorhizobium-Medicago model. Nature Reviews Microbiology. 5(8): 619-833. HTML
[14] Oldroyd GE, Murray JD, Poole PS, Downie JA. (2011) The rules of engagement in the legume-rhizobial symbiosis. Annual Review of Genetics. 45: 119-144. HTML
[15] Brewin NJ. (2004) Plant cell wall remodelling in the rhizobium-legume symbiosis. Critical Reviews in Plant Sciences. 23(4): 293-316. HTML
[16] Kereszt A, Mergaert P, Kondorosi É. (2011) Bacteroid develoment in legume nodules: evolution of mutual benefit or of sacrificial victims? Molecular Plant-Microbe Interactions. 24(11): 1300-1309. HTML
[17] Haag AF, Arnold MFF, Myka KK, Kersher B, Dall'Angelo S, Zanda M, Mergaert P, Ferguson GP. (2013) Molecular insights into bacteroid development during Rhizobium-legume symbiosis. FEMS Microbiology Reviews. 37(3): 364-383. HTML
[18] Roux B, Rodde N, Jardinaud MG, Timmers T, Sauviac L, Cottret L, Carrère S, Sallet E, Courcelle E, Moreau S, Debellé F, Capela D, de Carvalho-Niebel F, Gouzy J, Bruand C, Gamas P. 2014. An integrated analysis of plant and bacterial gene expression in symbiotic root nodules using laser-capture microdissection coupled to RNA sequencing. The Plant Journal. 77(6): 827-837. HTML
[19] Udvardi M, Poole PS. (2013) Transport and metabolism in legume-rhizobia symbioses. Annual Review of Plant Biology. 64: 781-805. HTML
[20] Vasse J, de Billy F, Camut S, Truchet G. (1990) Correlation between ultrastructural differentiation of bacteroids and nitrogen fixation in alfalfa nodules. Journal of Bacteriology. 172(8): 4295-4306. HTML
[21] diCenzo GC, Zamani M, Checcucci A, Fondi M, Griffitts JS, Finan TM, Mengoni A. (2019) Multi-disciplinary approaches for studying rhizobium – legume symbioses. Canadian Journal of Microbiology. 65: 1-33. HTML
[22] Guefrachi I, Verly C, Kondorisi É, Alunni B, Mergaert P. (2015) Role of the bacterial BacA ABC-transporter in chronic infection of nodule cells by Rhizobium bacteria. In Biological Nitrogen Fixation. Edited by F. J. de Bruin. Wiley Blackwell, Hobokon, New Jersy, USA. HTML
[23] diCenzo GC, Zamani M, Ludwig H, Finan TM. (2017) Heterologous complementation reveals a specialized role for BacA in the Medicago – Sinorhizobium meliloti symbiosis. Molecular Plant-Microbe Interactions. 30(4): 312-324. HTML
[24] Domenech P, Kobayashi H, LeVier K, Walker GC, Barry CE 3rd. (2009) BacA, and ABC transporter involved in maintenance of chronic murine infections with Mycobacterium tuberculosis. Journal of Bacteriology. 191(2): 477-485. HTML
[25] diCenzo GC, Finan TM. (2017) The divided bacterial genome: structure, function, and evolution. Microbiology and Molecular Biology Reviews. 81(3): e00019-17. HTML
[26] Harrison PW, Lower RP, Kim NK, Young JP (2010) Introducing the bacterial 'chromid': not a chromosome, not a plasmid. Trends in Microbiology. 18(4): 141-148. HTML
[27] Oresnik IJ, Liu SL, Yost CK, Hynes MF. (2000) Megaplasmid pRme2011a of Sinorhizobium meliloti is not required for viability. Journal of Bacteriology. 182(12): 3582-2586. HTML
[28] diCenzo GC, Muhammed Z, Østerås M, O’Brien SAP, Finan TM. (2017) PckR is a key regulator of the glycolytic and gluconeogenic central metabolic pathways in Sinorhizobium meliloti. Genetics. 207(3): 961-974. HTML
[29] diCenzo GC, Benedict A, Fondi M, Walker G, Finan TM, Mengoni A, Griffitts JS. (2018) Robustness encoded across essential and accessory replicons of the ecologically versatile bacterium Sinorhizobium meliloti. PLOS Genetics. 14(4): e1007357. HTML
[30] Mitsch MJ, diCenzo GC, Cowie A, Finan TM. (2018) Succinate transport is not essential for symbiotic nitrogen fixation by Sinorhizobium meliloti nor Rhizobium leguminosarum. Applied and Environmental Microbiology. 84(1): e01561-17. HTML
[31] Kuzmanović N, Fagorzi C, Mengoni A, Lassalle F, diCenzo GC. (2022) Taxonomy of Rhizobiaceae revisited: proposal of a new framework for genus delimitation. International Journal of Systematic and Evolutionary Microbiology. 72(3): 005243. HTML





